Activity-based Crosslinking to Identify Substrates of Thioredoxin-domain Proteinsin Malaria Parasites
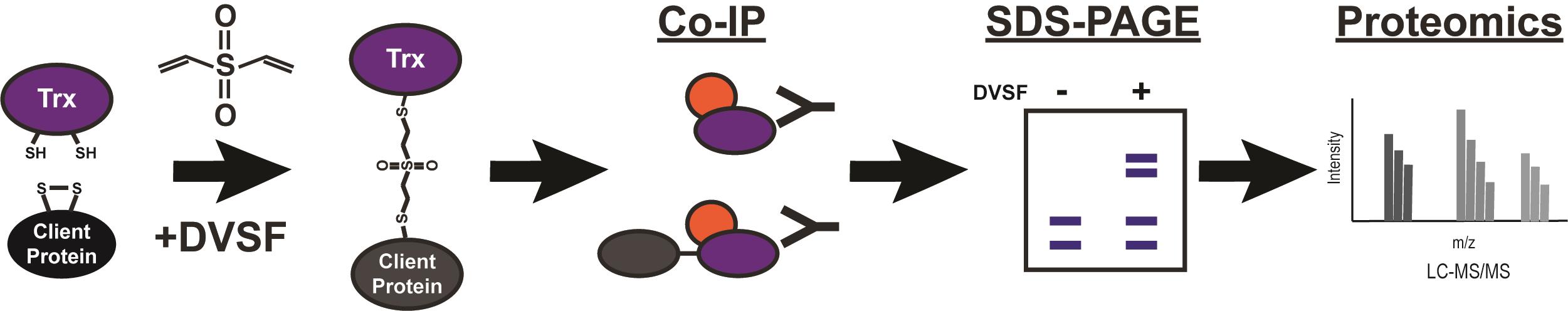
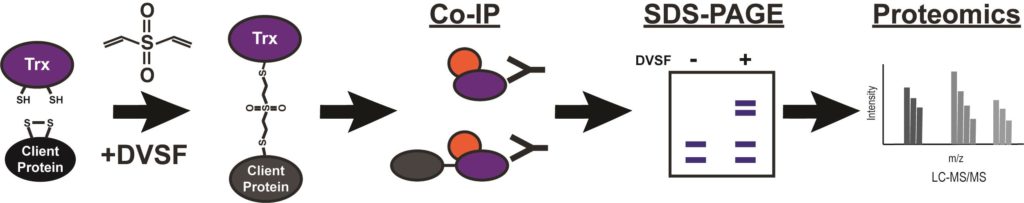 Malaria remains a major public health issue, infecting nearly 220 million people every year. The spread of drug-resistant strains of Plasmodium falciparum around the world threatens the progress made against this disease. Therefore, identifying druggable and essential pathways in P. falciparum parasites remains a major area of research. One poorly understood area of parasite biology is the formation of disulfide bonds, which is an essential requirement for the folding of numerous proteins. Specialized chaperones with thioredoxin (Trx) domains catalyze the redox functions necessary for breaking incorrect and forming correct disulfide bonds in proteins. Defining the substrates of these redox chaperones is difficult and immunoprecipitation based assays cannot distinguish between substrates and interacting partners. Further, the substrate or client interactions with the redox chaperones are usually transient in nature. Activity based crosslinkers that rely on the nucleophilic cysteines on Trx domains and the disulfide bond forming cysteines on clients provide an easily scalable method to trap and identify the substrates of Trx-domain containing chaperones. The cell permeable crosslinker divinyl sulfone (DVSF) is active only in the presence of nucleophilic cysteines in proteins and, therefore, traps Trx domains with their substrates, as they form mixed disulfide bonds during the course of their catalytic activity. This allows the identification of substrates that rely on Trx activity for their folding, as well as discovering small molecules that interfere with Trx domain activity. Graphic abstract: Identification of thioredoxin domain substrates via divinylsulfone crosslinking and immunoprecipitation-mass spectrometry.
Malaria remains a major public health issue, infecting nearly 220 million people every year. The spread of drug-resistant strains of Plasmodium falciparum around the world threatens the progress made against this disease. Therefore, identifying druggable and essential pathways in P. falciparum parasites remains a major area of research. One poorly understood area of parasite biology is the formation of disulfide bonds, which is an essential requirement for the folding of numerous proteins. Specialized chaperones with thioredoxin (Trx) domains catalyze the redox functions necessary for breaking incorrect and forming correct disulfide bonds in proteins. Defining the substrates of these redox chaperones is difficult and immunoprecipitation based assays cannot distinguish between substrates and interacting partners. Further, the substrate or client interactions with the redox chaperones are usually transient in nature. Activity based crosslinkers that rely on the nucleophilic cysteines on Trx domains and the disulfide bond forming cysteines on clients provide an easily scalable method to trap and identify the substrates of Trx-domain containing chaperones. The cell permeable crosslinker divinyl sulfone (DVSF) is active only in the presence of nucleophilic cysteines in proteins and, therefore, traps Trx domains with their substrates, as they form mixed disulfide bonds during the course of their catalytic activity. This allows the identification of substrates that rely on Trx activity for their folding, as well as discovering small molecules that interfere with Trx domain activity. Graphic abstract: Identification of thioredoxin domain substrates via divinylsulfone crosslinking and immunoprecipitation-mass spectrometry.
David W Cobb, Grace S Woods, Vasant Muralidharan. Bio Protoc. 2022 Feb 20;12(4):e4322. doi: 10.21769/BioProtoc.4322.