VEuPathDB: the eukaryotic pathogen, vector and host bioinformatics resource center in 2023
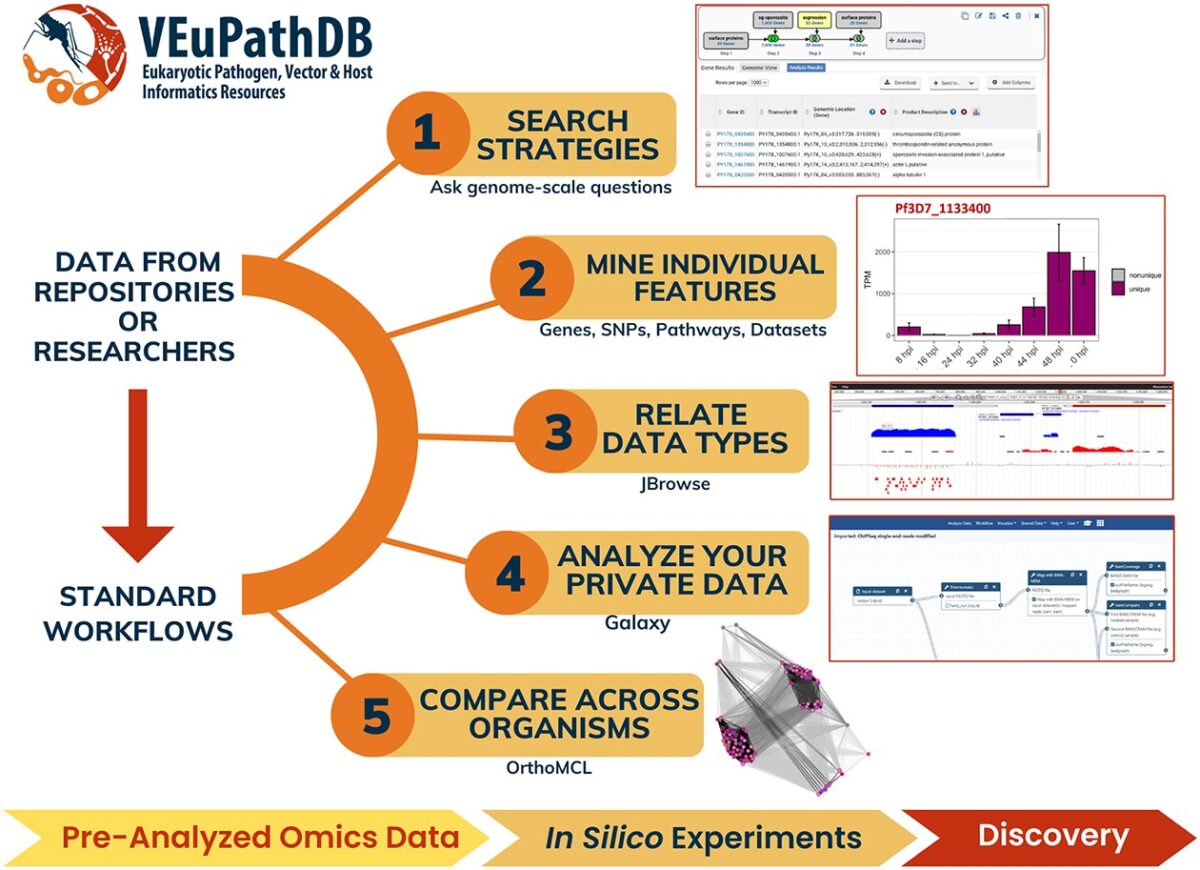
The Eukaryotic Pathogen, Vector and Host Informatics Resource (VEuPathDB, https://veupathdb.org) is a Bioinformatics Resource Center funded by the National Institutes of Health with additional funding from the Wellcome Trust. VEuPathDB supports >600 organisms that comprise invertebrate vectors, eukaryotic pathogens (protists and fungi) and relevant free-living or non-pathogenic species or hosts. Since 2004, VEuPathDB has analyzed …