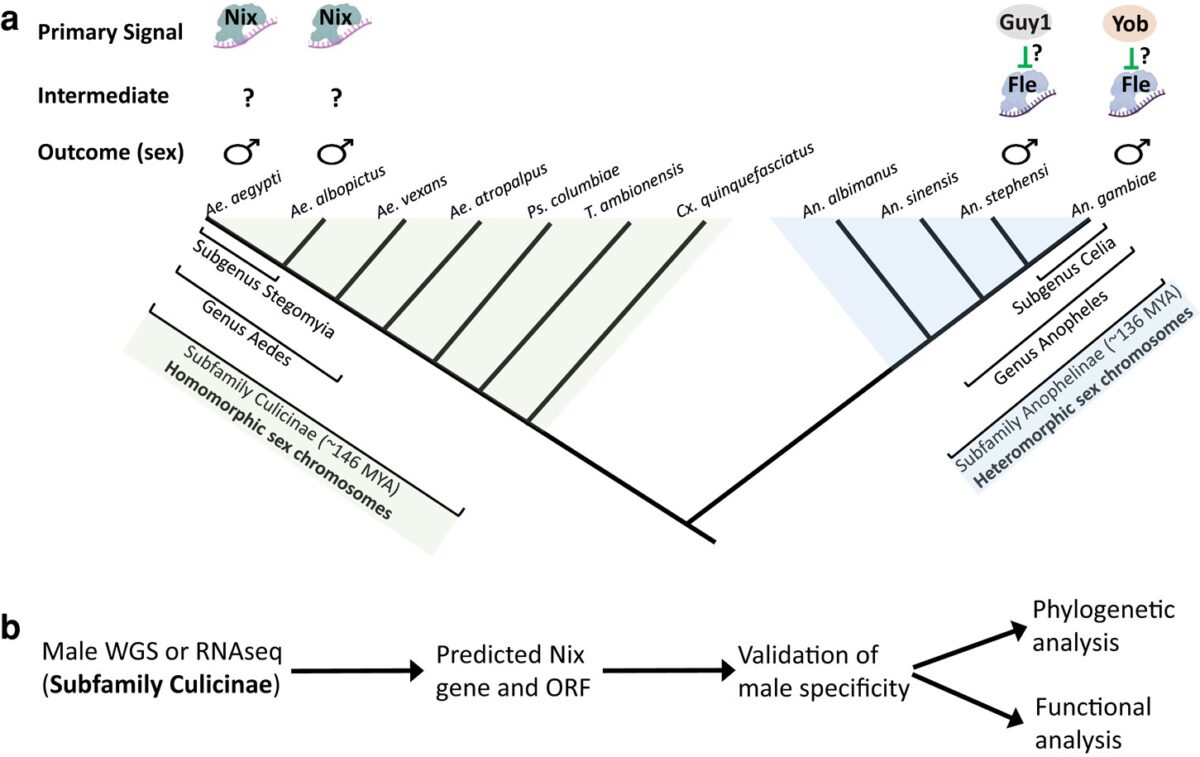
On the origin and evolution of the mosquito male-determining factor Nix
The mosquito family Culicidae is divided into two subfamilies named the Culicinae and Anophelinae. Nix, the dominant male-determining factor, has only been found in the culicines Aedes aegypti and Ae. albopictus, two important arboviral vectors that belong to the subgenus Stegomyia. Here we performed sex-specific whole-genome sequencing and RNAseq of divergent mosquito species and explored …