Researchers discover malaria gene needed to make pair of invasion organelles
Plasmodium RON11 triggers biogenesis of the merozoite rhoptry pair and is essential for erythrocyte invasion
Malaria is a global and deadly human disease caused by the apicomplexan parasites of the genus Plasmodium. Parasite proliferation within human red blood cells (RBCs) is associated with the clinical manifestations of the disease. This asexual expansion within human RBCs begins with the invasion of RBCs by P. falciparum, which is mediated by the …
Trainee Spotlight: Grace Vick
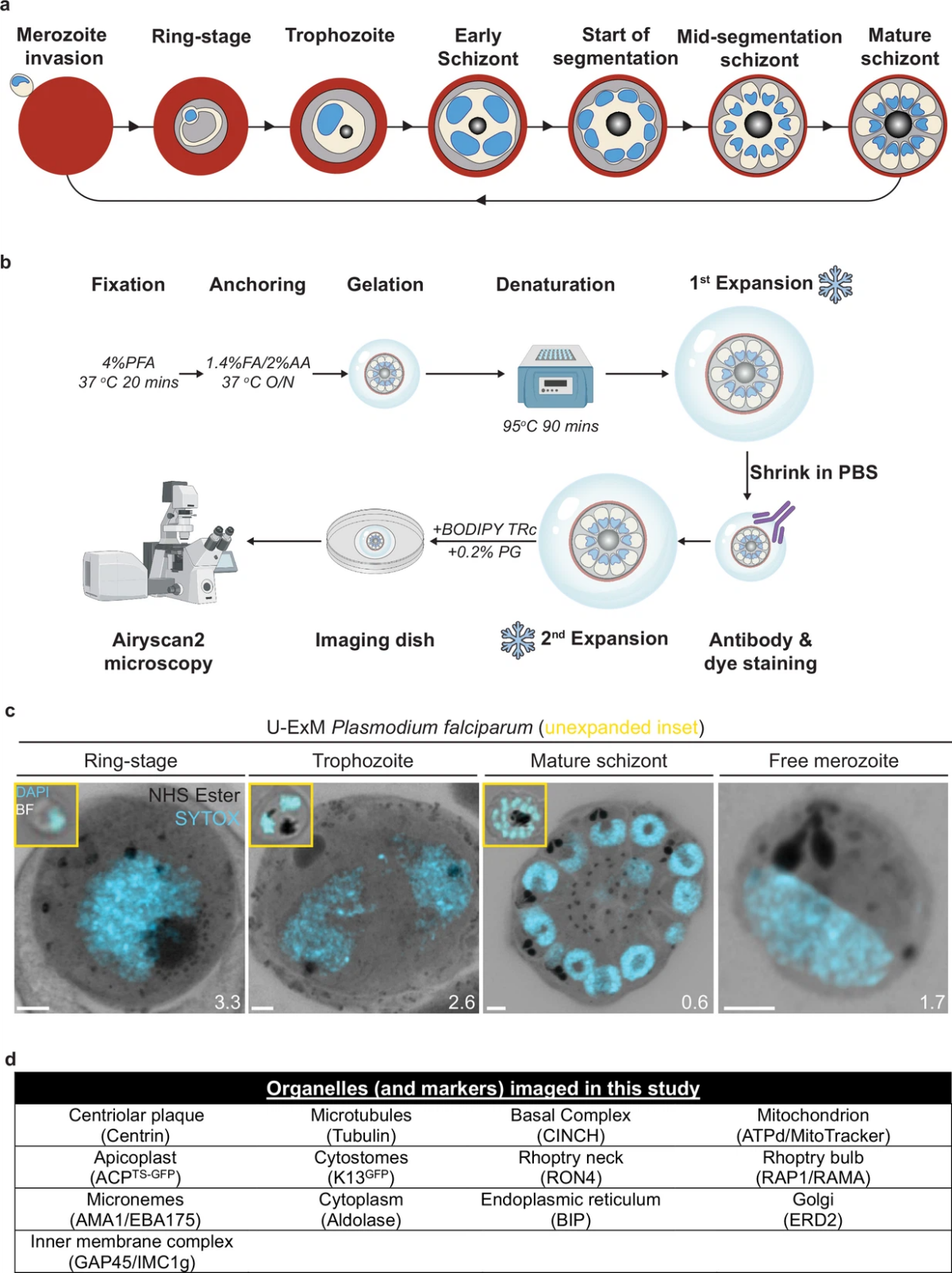
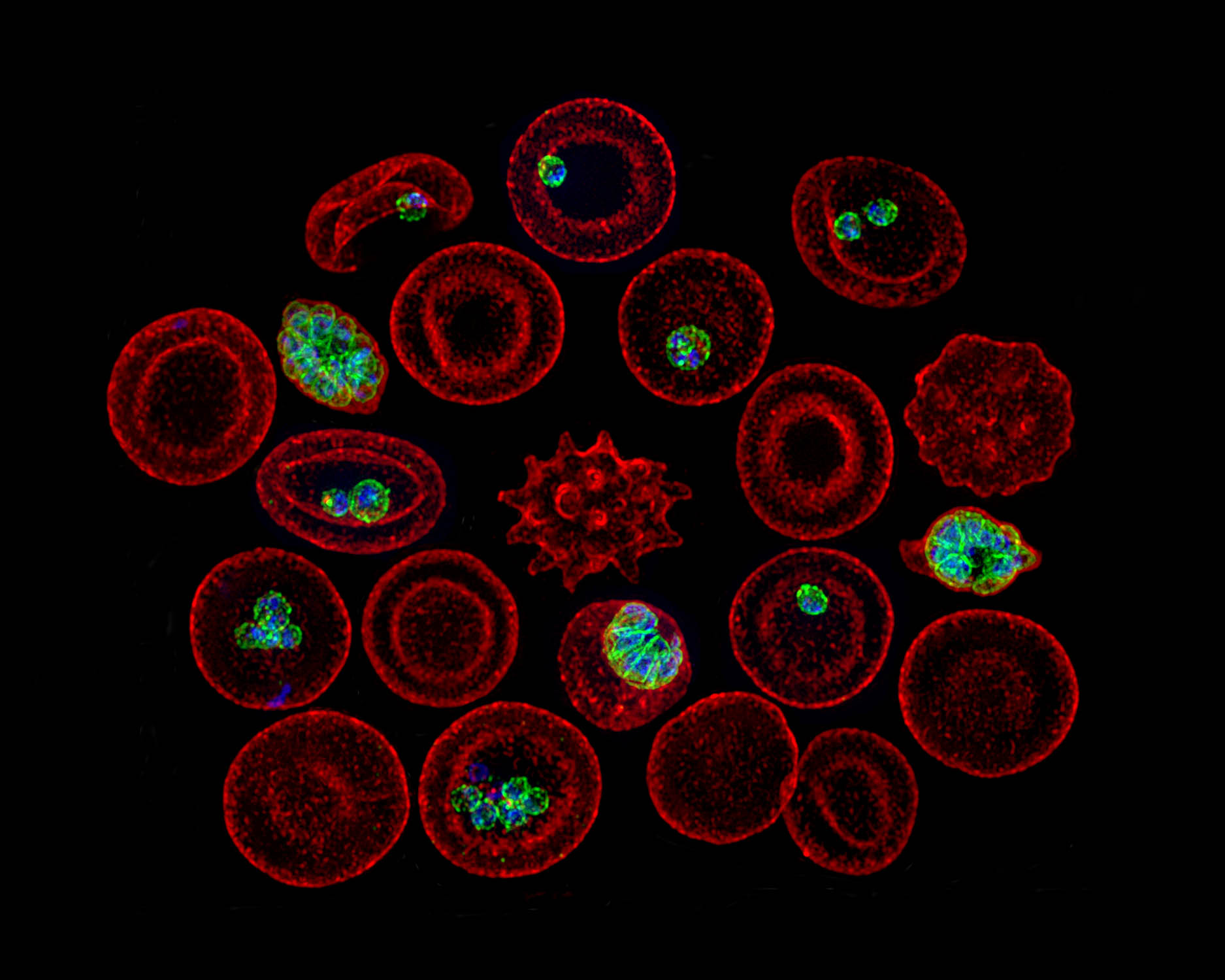
Atlas of Plasmodium falciparum intraerythrocytic development using expansion microscopy
Apicomplexan parasites exhibit tremendous diversity in much of their fundamental cell biology, but study of these organisms using light microscopy is often hindered by their small size. Ultrastructural expansion microscopy (U-ExM) is a microscopy preparation method that physically expands the sample by ~4.5×. Here, we apply U-ExM to the human malaria parasite Plasmodium falciparum during the asexual …
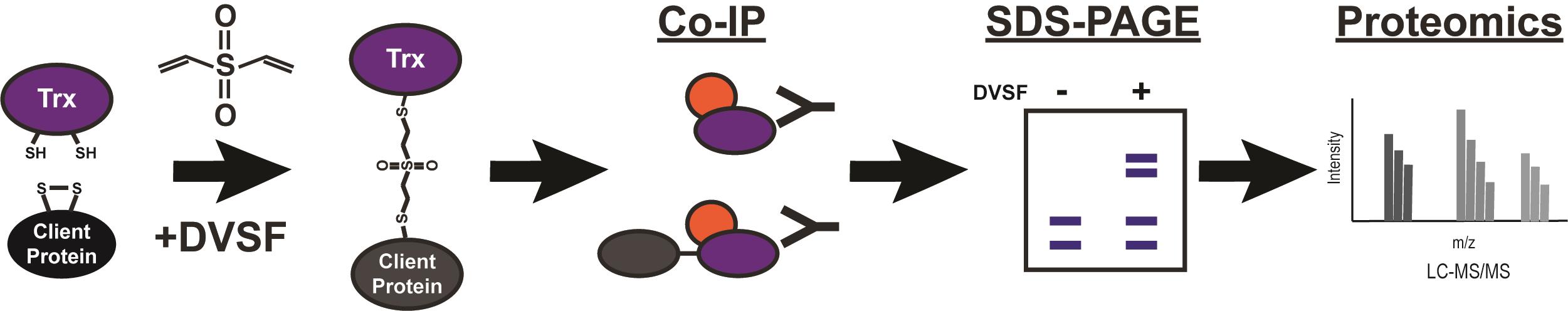
Time-resolved proximity biotinylation implicates a porin protein in export of transmembrane malaria parasite effectors
The malaria-causing parasite, Plasmodium falciparum completely remodels its host red blood cell (RBC) through the export of several hundred parasite proteins, including transmembrane proteins, across multiple membranes to the RBC. However, the process by which these exported membrane proteins are extracted from the parasite plasma membrane for export remains unknown. To address this question, we …