Pseudokinase NRP1 facilitates endocytosis of transferrin in the African trypanosome
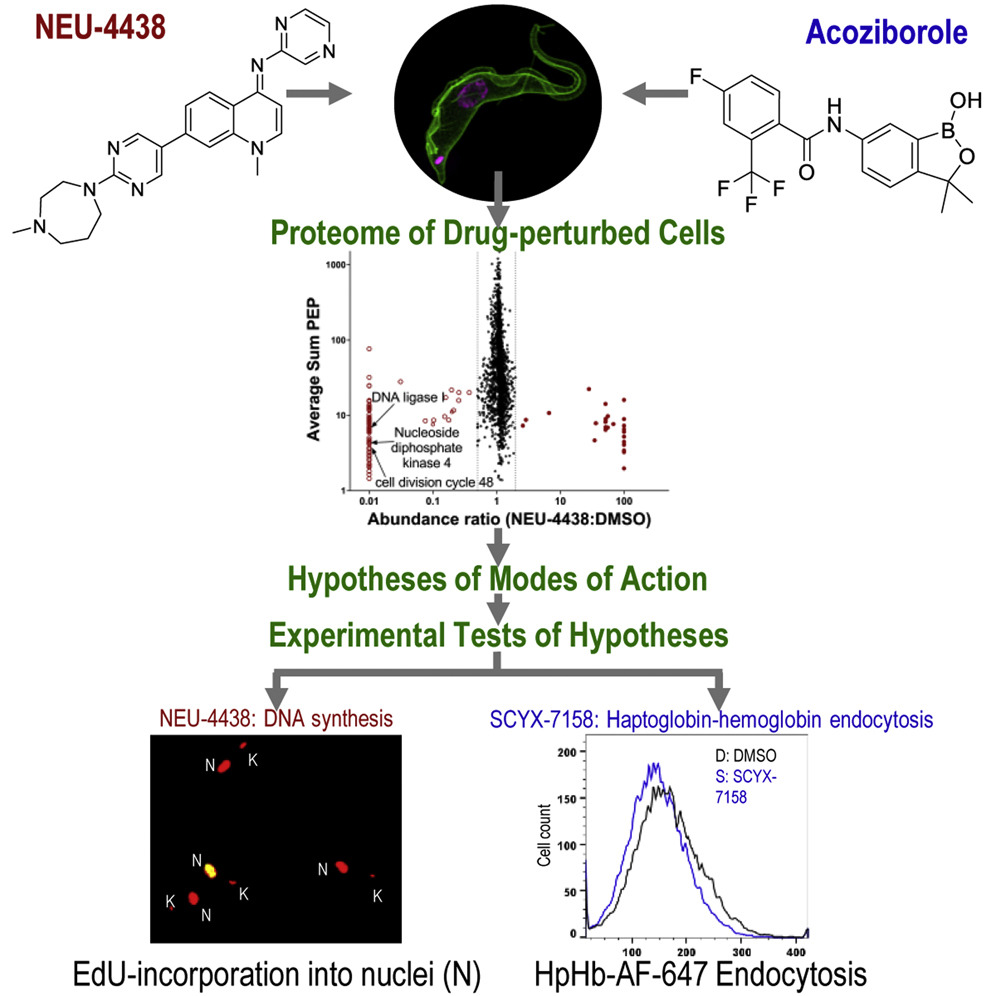
Hypothesis-generating proteome perturbation to identify NEU-4438 and acoziborole modes of action in the African Trypanosome
Casein kinase TbCK1.2 regulates division of kinetoplast DNA, and movement of basal bodies in the African trypanosome
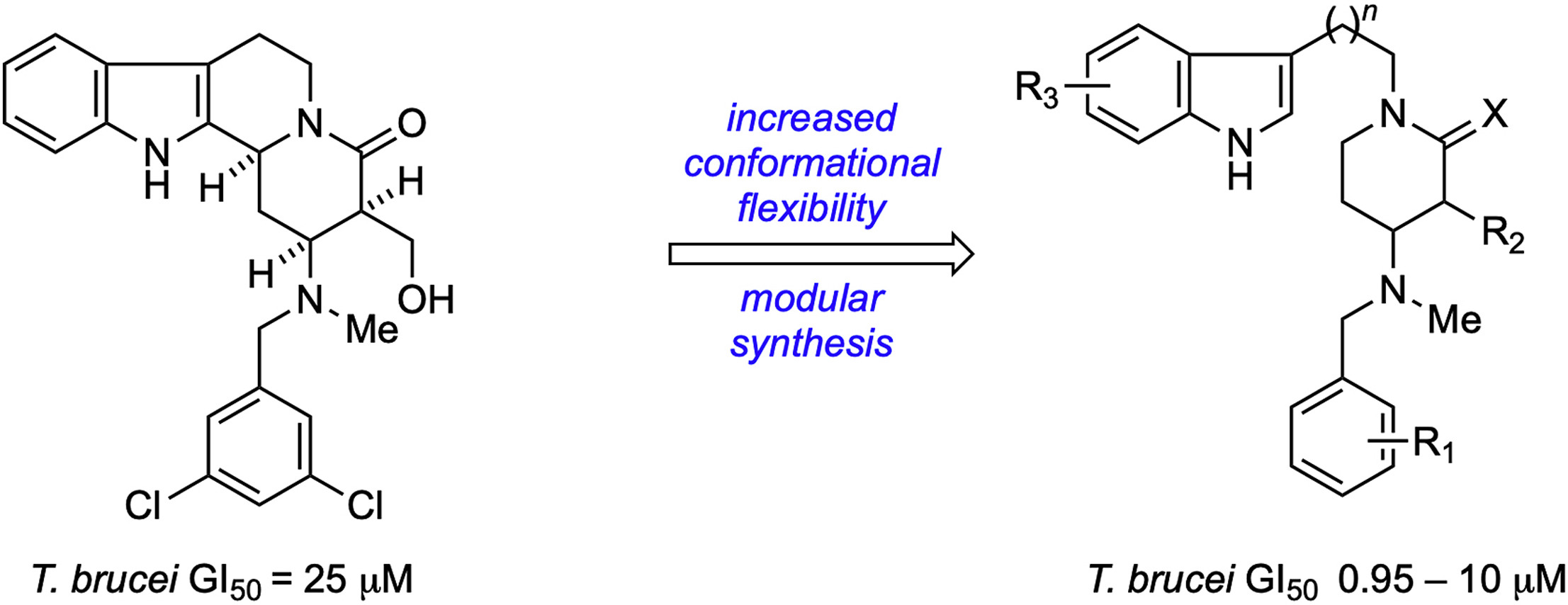
Design, Synthesis, and Evaluation of Novel Anti-Trypanosomal Compounds
Kinetoplast Division Factors in a Trypanosome
Anilinoquinoline based inhibitors of trypanosomatid proliferation
Series of Alkynyl-Substituted Thienopyrimidines as Inhibitors of Protozoan Parasite Proliferation
Trainee Adds New Tool to the Trypanosome Toolbox
When Ph.D. trainee Justin Wiedeman started investigating the role of protein kinase TbCK1.2, an enzyme found near the flagellum of Trypanosoma brucei, he quickly ran into a problem common to parasitologists. He needed a better tool for visualizing the membranes of this parasite. Since none of the membrane probes on the market quite did the …
New UGA Drug Discovery Core lab works to develop treatment of leading diseases
Athens, Ga. – The University of Georgia has created the Drug Discovery Core laboratory, a campus-wide collaborative facility designed to hasten the development of therapeutic drugs for a number of major diseases. A survey distributed to UGA researchers in 2016 identified chemical screening and toxicity profiling as the most critical needs for enhancing drug discovery …