Hypothesis-generating proteome perturbation to identify NEU-4438 and acoziborole modes of action in the African Trypanosome
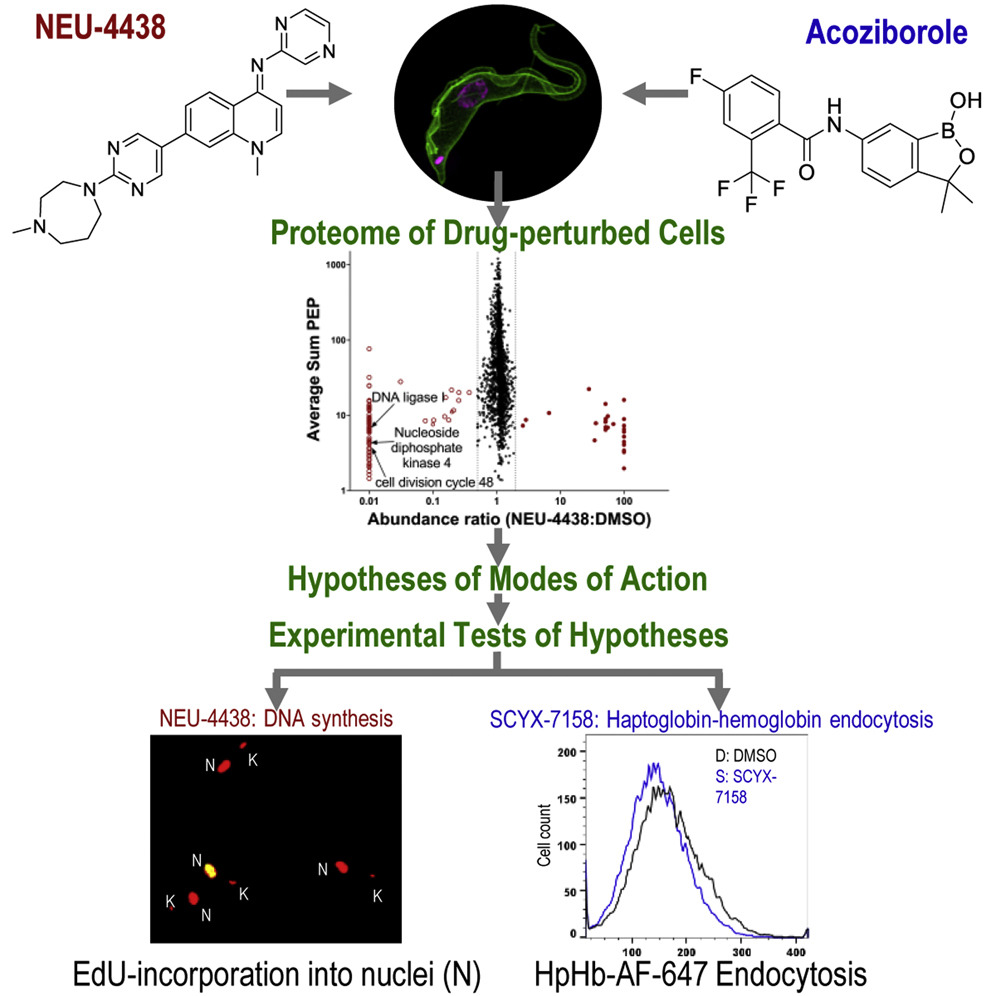
NEU-4438 is a lead for the development of drugs against Trypanosoma brucei, which causes human African trypanosomiasis. Optimized with phenotypic screening, targets of NEU-4438 are unknown. Herein, we present a cell perturbome workflow that compares NEU-4438’s molecular modes of action to those of SCYX-7158 (acoziborole). Following a 6 h perturbation of trypanosomes, NEU-4438 and acoziborole reduced steady-state amounts of 68 and 92 unique proteins, respectively. After analysis of proteomes, hypotheses formulated for modes of action were tested: Acoziborole and NEU-4438 have different modes of action. Whereas NEU-4438 prevented DNA biosynthesis and basal body maturation, acoziborole destabilized CPSF3 and other proteins, inhibited polypeptide translation, and reduced endocytosis of haptoglobin-hemoglobin. These data point to CPSF3-independent modes of action for acoziborole. In case of polypharmacology, the cell-perturbome workflow elucidates modes of action because it is target-agnostic. Finally, the workflow can be used in any cell that is amenable to proteomic and molecular biology experiments.
Amrita Sharma, Michael Cipriano, Lori Ferrins, Stephen L Hajduk, Kojo Mensa-Wilmot. iScience. 2022 Oct 7;25(11):105302. doi: 10.1016/j.isci.2022.105302. eCollection 2022 Nov 18.
