Melissa Sleda, a Ph.D. trainee is Silvia Moreno’s laboratory, is in her third year at UGA. She is originally from Sandusky, Michigan and attended Lawrence Technological University where she majored in Molecular and Cell Biology with a minor in Chemistry. At UGA, she has held positions as the Secretary for the Cell Bio Grad …
Trainee Spotlight: Alona Botnar
T32 trainee Alona Botnar is entering her fifth year as a Ph.D. candidate in Dr. Dennis Kyle’s laboratory. She is from Doylestown, Pennsylvania and completed her B.S. in Chemistry with a minor in Biochemistry at the University of Georgia in 2015. During her undergraduate career, she also worked at Janssen, a pharmaceutical company of Johnson …
Age influences thermal tolerance in Asian malaria mosquito
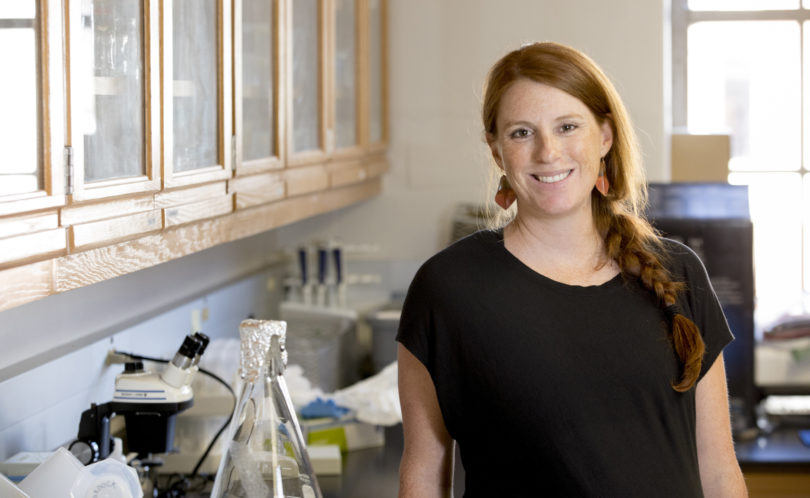
Environmental portrait of Courtney Murdock in her laboratory at UGA's School of Veterinary Medicine (Dorothy Kozlowski) Malaria disease transmission models are important tools for controlling and eliminating disease spread. However, a model is only as good as the assumptions about the various variables. Dr. Courtney Murdock, a member of the UGA’s Center for Tropical and …
NIH awards CTEGD $1.9 million to support training in tropical and emerging global diseases
UGA’s Center for Tropical and Emerging Global Diseases has been awarded $1.9 million from the National Institutes of Health to continue its pre- and post-doctoral training program for the next five years. First funded in 2004, CTEGD has received nearly $2 million from NIH to train the next generation of scientists in the fight against …
Trainee Spotlight: Emma Troth
Emma Troth, a Ph.D. trainee in Dennis Kyle's laboratory, is entering her fourth year at UGA. She is originally from Eureka, Illinois, and attended Bradley University where she majored in Biology with a minor in Ethics. While at UGA, Emma has served as president of the CTEGD Graduate Student Association (2019-2020) and is currently the …
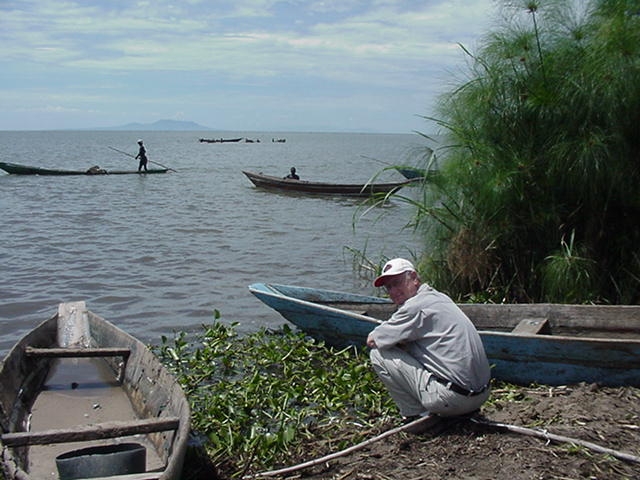
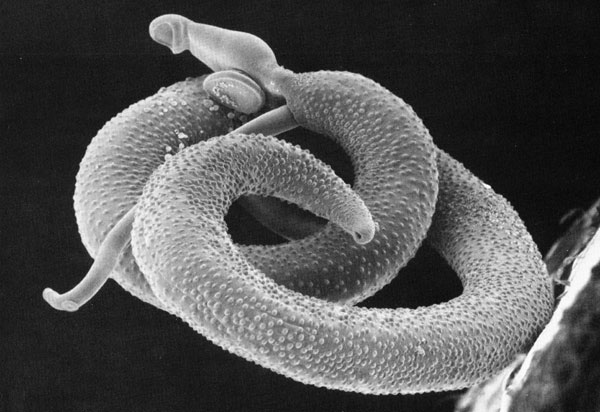
SCORE – a decade of operational research with a lasting legacy
It all began with a simple phone call and now, more than a decade later, the Schistosomiasis Consortium for Operational Control and Evaluation (SCORE) is preparing to pass the baton to new groups of investigators working on understanding and controlling schistosomiasis. Under the direction of Dan Colley, a member of the Center for Tropical and …