EdU Incorporation To Assess Cell Proliferation and Drug Susceptibility in Naegleria fowleri
Characterization of the Tubovesicular Network in Plasmodium vivax Liver Stage Hypnozoites and Schizonts
Challenges for Cryptosporidium Population Studies
Calcium signaling through a Transient Receptor Channel is important for Toxoplasma gondii growth
Diet-Microbiota Interactions Alter Mosquito Development
Gut microbes and diet can both strongly affect the biology of multicellular animals, but it is often difficult to disentangle microbiota-diet interactions due to the complex microbial communities many animals harbor and the nutritionally variable diets they consume. While theoretical and empirical studies indicate that greater microbiota diversity is beneficial for many animal hosts, there …
Dennis Kyle Featured Guest on People, Parasites & Plagues Podcast
Trypanosoma cruzi Letm1 is involved in mitochondrial Ca 2+ transport, and is essential for replication, differentiation, and host cell invasion
Lacto-N-fucopentaose-III ameliorates acute and persisting hippocampal synaptic plasticity and transmission deficits in a Gulf War Illness mouse model
Aims: The present study investigated if treatment with the immunotherapeutic, lacto-N-fucopentaose-III (LNFPIII), resulted in amelioration of acute and persisting deficits in synaptic plasticity and transmission as well as trophic factor expression along the hippocampal dorsoventral axis in a mouse model of Gulf War Illness (GWI). Main methods: Mice received either coadministered or delayed LNFPIII treatment throughout or …
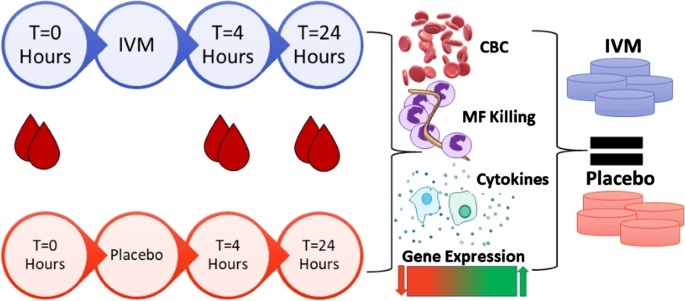
Lack of detectable short-term effects of a single dose of ivermectin on the human immune system
Background: Ivermectin is widely used in human and animal medicine to treat and prevent parasite nematode infections. It has been suggested that its mode of action requires the host immune system, as it is difficult to reproduce its clinical efficacy in vitro. We therefore studied the effects of a single dose of ivermectin (Stromectol®-0.15 mg/kg) on …